Single-cell sequencing facts for kids
Single-cell sequencing looks at the genetic information, like DNA or RNA, from one single cell at a time. This is different from older methods that looked at millions of cells all mixed together. By studying individual cells, scientists can see tiny differences between them. This helps us better understand how cells work, especially in diseases like cancer or during development. For example, in cancer, looking at the DNA of single cells can show tiny changes (mutations) in small groups of cells. In development, studying the RNA from individual cells can help us understand how different cell types appear and behave.
Contents
What is Single-Cell Sequencing?
A typical human cell holds about 3.3 billion pairs of DNA "letters" and 600 million RNA "messages." Usually, older methods like Sanger sequencing or next generation sequencing use a mix of millions of cells. By looking closely at the DNA and RNA from just one cell, scientists can learn a lot about how that cell works.
Single-cell sequencing usually involves these steps:
- First, a single cell is carefully separated.
- Then, its DNA or RNA is taken out and copied many times.
- Next, the copied material is prepared for sequencing.
- After that, the actual sequencing happens.
- Finally, computers analyze all the data.
It is harder to do single-cell sequencing than to sequence from many cells at once. A single cell has very little starting material. This means it is easy for the material to break down, get lost, or become contaminated. Also, because there is so little DNA or RNA, scientists often need to make many copies. This copying process can sometimes lead to uneven results or errors.
Despite these challenges, new improvements have made single-cell sequencing a powerful tool. It helps scientists study complex samples, rare cell types, and how cells are related. It can also help understand diseases and tiny living things that cannot be grown in a lab. Because of its importance, Nature Publishing Group chose single-cell sequencing as the "Method of the Year" in 2013.
Looking at a Cell's DNA: Genome Sequencing
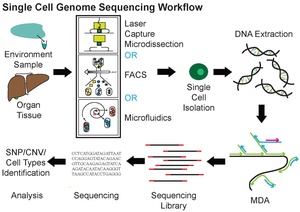
Single-cell DNA genome sequencing means taking one cell, making many copies of its entire DNA or a specific part, and then using next generation sequencing machines to read it. This method helps scientists study normal body functions and diseases in animals. Looking at individual cells can show how tiny genetic differences within a tumor affect cancer growth or how it responds to treatment.
In the world of tiny living things (microbes), the DNA from a single-celled organism is called a "single amplified genome" (SAG). Advances in this sequencing have allowed scientists to get genetic information from microbes that are hard to grow in a lab. Even though these SAGs might not be complete, new computer tools can help put together almost full genomes from them.
How Scientists Study DNA in Single Cells
One common way to copy DNA from a single cell is called Multiple Displacement Amplification (MDA). This method can turn tiny amounts of DNA into much larger amounts needed for sequencing. MDA uses special ingredients, including random starting pieces (primers) and an enzyme called DNA polymerase. In a warm, steady temperature, the DNA is copied many times. As new DNA strands are made, they push aside older ones, creating many copies from each original DNA piece. MDA products can be quite long, making them useful for DNA sequencing. A newer version, called WGA-X, improved this method, especially for DNA with lots of G and C letters.
Another method is called MALBAC. Like MDA, it starts by copying DNA at a steady temperature. However, MALBAC uses special primers that help the newly copied DNA form "loops." These loops stop the DNA from being copied too much, which helps create more even copies across the entire genome. Later, these loops are opened up, and the DNA is copied again using a method called PCR.
When comparing MDA and MALBAC, MDA generally covers more of the genome. However, MALBAC provides more even coverage across the DNA. MDA might be better for finding tiny single-letter changes (SNPs). MALBAC is often preferred for finding larger changes in the number of DNA copies.
A method called Strand-seq is good for finding big changes in the structure of DNA. It can find changes that are at least 200,000 DNA letters long. Strand-seq is less likely to have certain errors that other methods might have. The best method to choose depends on what scientists want to find.
Challenges When Sequencing DNA
When MDA is used to copy DNA from individual cells, the copies can be very uneven. This means some parts of the DNA are copied too much, and others are copied too little or even missed. This can lead to losing some important DNA sequences. Also, MDA sometimes struggles with DNA regions that have a high percentage of G and C letters.
Finding tiny single-letter changes (SNPs) and changes in the number of DNA copies (CNVs) can be hard in single-cell sequencing. This is because there is so little DNA from one cell. Even after copying, the coverage might be low, leading to errors. MDA can also miss one of the two copies of a gene in a cell. Scientists are working on new computer programs to help find true changes and remove false ones.
Strand-seq helps overcome some of these problems for finding big DNA changes. It can find many types of structural changes, including complex ones. However, Strand-seq needs cells that are dividing to work. It also cannot find changes smaller than 200,000 DNA letters.
What We Learn from Single-Cell DNA Sequencing
Microbes: Single-cell DNA sequencing is very useful for studying microbes (like bacteria). Many microbes are hard to grow in a lab. This method allows scientists to get their DNA sequences without needing to grow them. This approach has been used in marine, soil, and other environments. It helps answer questions about how microbes live, change over time, and their potential for medicine or technology.
Cancer: This sequencing is also becoming important for studying cancer. Scientists can analyze fresh or frozen tumor samples. They can look for changes in DNA copy numbers, single-letter changes, and rearrangements. Single-cell DNA sequencing is especially good at showing how complex and varied cancer cells can be within a single tumor. It helps reveal how different mutations might work together in individual cancer cells. This understanding can lead to new ways to fight cancer.
Exploring Cell Control: DNA Methylation
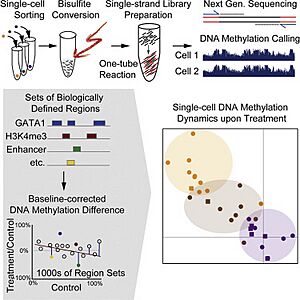
Single-cell DNA methylome sequencing measures DNA methylation. DNA methylation is like a chemical tag on DNA. It can turn genes "on" or "off" without changing the actual DNA sequence. In animals, a common type of methylation (5mC) is found throughout the DNA. It plays a big role in controlling which genes are active. By sequencing 5mC in individual cells, scientists can see how these "epigenetic" changes make genetically identical cells behave differently.
How Scientists Find Methylation Marks
Bisulfite sequencing is the main method for finding 5mC in single cells. This method treats DNA with a chemical called bisulfite. Bisulfite changes normal cytosine letters into uracil, but it leaves methylated cytosine letters (5mC) untouched. So, after this treatment, only the methylated cytosines remain. Scientists then compare this treated sequence to the original DNA to find where the methylation marks are.
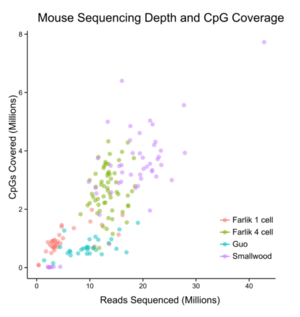
In 2014, scientists were able to do whole-genome bisulfite sequencing on single cells. This method improved how DNA is prepared, allowing more DNA fragments to be copied. Using deep sequencing, this method can find about 40% of the important CpG sites in each cell. It is important that DNA cannot be copied before bisulfite treatment, because the methylation marks would not be copied correctly.
Another method is called single-cell reduced representation bisulfite sequencing (scRRBS). This method focuses on areas of the DNA that are rich in CpG sites, where methylation often happens. This makes sequencing cheaper than looking at the whole genome. However, it also means less of the genome is covered. When this method is used on many cells at once, it finds most CpG sites in gene promoters. In single cells, about 40% of these sites are found. Scientists can also combine a few single cells to increase the coverage.
Difficulties in Studying Methylation
While bisulfite sequencing is widely used, the chemical treatment is harsh. It can break and damage the DNA. This problem becomes even bigger when working with the tiny amount of DNA from a single cell. Other methods to find DNA methylation exist, such as using special enzymes.
Nanopore sequencing is a newer method that can directly read methylation without breaking or changing the DNA. Nanopore sequencing has been used to study methylation in bacteria. However, this technique has not yet been made small enough to work on single cells.
Why Studying Methylation is Important
Single-cell DNA methylation sequencing has been used to find epigenetic differences in cells that have the same DNA. For example, scientists successfully sorted mixed cell populations into distinct types based on their methylation patterns. This method also helps study single cells during early development. It shows how different cell types emerge from a single embryo. Single-cell whole-genome bisulfite sequencing has also been used to study rare but active cancer cells, such as circulating tumor cells.
Understanding Gene Activity: RNA Sequencing
Single-cell RNA sequencing (scRNA-seq) shows which genes are active in individual cells. It is considered the best way to understand what a cell is doing and what type of cell it is. Even though it is impossible to get every piece of RNA information from each cell, scientists can find patterns of gene activity. This can help discover rare cell types that were never seen before. For example, scientists using scRNA-seq on a type of cancer called neuroblastoma found a rare cancer cell. This discovery could lead to new ways to treat the disease.
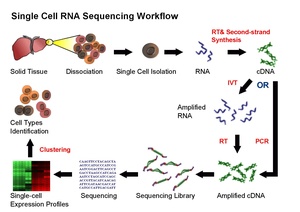
How Scientists Sequence RNA in Single Cells
Current scRNA-seq methods involve separating single cells and their RNA. Then, they follow similar steps to sequencing RNA from many cells:
- Reverse Transcription (RT): RNA is turned into DNA copies (called cDNA).
- Amplification: The cDNA is copied many times.
- Library Generation: The copied cDNA is prepared for sequencing.
- Sequencing: The DNA sequences are read.
Early methods put individual cells into separate containers. More recent methods put single cells into tiny oil droplets using special devices. Inside each droplet, the RNA is turned into cDNA. Each droplet also has a unique DNA "barcode." This barcode labels all the cDNA that came from that single cell. After this step, cDNA from many cells can be mixed together for sequencing. The unique barcode helps scientists know which RNA came from which cell.
A big challenge for scRNA-seq is making sure the original amounts of RNA in a cell are kept accurate. Also, it's hard to find rare RNA messages. The reverse transcription step is very important. How well it works determines how much of the cell's RNA will be analyzed.
Example of a Droplet-Based Platform: The 10X Method
This platform allows scientists to study gene activity in many individual cells at once. It uses tiny oil droplets to capture single cells and prepare them for sequencing.
Here's how it generally works:
- First, individual cells are captured separately and broken open.
- Then, the RNA messages (mRNA) are turned into cDNA. Special starting pieces (primers) that attach to a "poly(A) tail" on mRNA are used.
- The copied cDNA is then used to create a library for sequencing.
The first step involves putting single cells into tiny gel beads inside oil droplets (called GEMs). An automated machine uses a special chip to combine cells, gel beads, and other ingredients with oil. Each GEM usually contains one cell, one gel bead, and chemicals for the RT reaction. On each gel bead are special DNA pieces with four parts: a piece for sequencing, a unique 10X barcode, a Unique Molecular Identifier (UMI), and a piece that captures mRNA.
Inside each GEM, the cell breaks open, and its RNA is turned into cDNA. All the cDNA from the same cell gets the same 10X barcode. The number of UMIs helps scientists count how much of each gene is active. This data is often used to classify cell types or find new groups of cells.
Finally, the prepared libraries are sequenced using a method like Illumina dye sequencing. This method reads the DNA by adding colored "letters" one by one. To read both the RNA sequences and the barcodes, special sequencing machines are needed.
This droplet-based platform can find rare cell types because it can process many cells (500 to 20,000 cells per sample). The process is easy and can recover up to 65% of the cells. The whole process takes about 8 hours, which is faster than some other methods. However, it needs fresh samples and only detects about 10% of the total mRNA.
The main difference between droplet-based and microwell-based methods is how they separate the cells.
Challenges in RNA Sequencing
Most RNA-seq methods rely on capturing a "poly(A) tail" to find mRNA and remove other common RNA types. This means they often miss other important RNA molecules that don't have this tail, like some long non-coding RNAs or microRNAs. Scientists are developing new methods to capture these other RNA types. For example, one method uses special "masks" to block common RNA molecules and focuses on small RNAs. Another method uses random starting pieces to capture all types of RNA, but it needs to be designed carefully for each species.
Bacteria and other tiny single-celled organisms (prokaryotes) are currently hard to study with single-cell RNA-seq. This is because their mRNA usually does not have a poly(A) tail. Also, the total amount of RNA in a single bacterium is very small. These two factors make it difficult to use scRNA-seq for bacteria.
Another limitation is that scRNA-seq only captures a "snapshot" of what a cell is doing at one moment. For large, branched cells like neurons in the brain, isolating the cell might only capture RNA from the main cell body. It is estimated that over 40% of the RNA in the brain is in the long branches of cells, which might be missed by current scRNA-seq methods.
What We Learn from Single-Cell RNA Sequencing
scRNA-seq is now widely used in many areas of biology. These include developmental biology, neurology, cancer research, immunology, and infectious disease studies.
Scientists have used computer methods to improve scRNA-seq data. They use information from large cancer datasets to build networks of genes that work together. Applying these networks to single-cell data helps them better detect mutations in individual cells.
Some scRNA-seq methods have also been used for single-celled microbes. For example, it has been used to study single malaria parasites or yeast cells. One study on yeast looked at how different yeast cells, even if genetically identical, reacted differently to salt stress. This showed that cells in a population can vary in their responses, which might help some survive in changing environments.
The first single-cell RNA study in a prokaryotic species used a special enzyme to remove common RNA and then copied the mRNA. This method showed good results but has not yet been tested with next-generation sequencing. Single-cell RNA-seq for bacteria would be very helpful for studying microbiomes. It would help find species that are rare and understand the differences among cell populations.
scRNA-seq has given us great insights into how embryos and organisms develop. This includes animals like the worm Caenorhabditis elegans, the flatworm Schmidtea mediterranea, and the axolotl Ambystoma mexicanum. The first vertebrate animals mapped this way were Zebrafish and Xenopus laevis. By studying many stages of their embryos, scientists could map the entire development process cell by cell. Science recognized these advances as the "Breakthrough of the Year" in 2018.
Getting Cells Ready: Important Steps
Separating Single Cells
There are several ways to get individual cells before copying and sequencing their DNA or RNA.
- Fluorescence-activated cell sorting (FACS) is a common method. It uses lasers to sort cells based on their properties.
- Individual cells can also be collected by micromanipulation. This involves using tiny tools, like a fine pipette, to pick up a single cell. This method is easy and low-cost but can be slow. It also relies on scientists correctly identifying cell types under a microscope.
- Laser-capture microdissection (LCM) can also be used. This method helps keep track of where a cell was located in a tissue. However, it can be hard to capture a whole single cell without also picking up material from nearby cells.
- Microfluidics are high-tech methods for isolating single cells. Both FACS and microfluidics are accurate and can isolate samples automatically. However, both methods require detaching cells from their natural environment first. This can sometimes change how the cells' genes are expressed.
How Many Cells Should We Study?
The different scRNA-seq methods vary in how well they capture RNA. This leads to differences in how many RNA messages are found from each single cell. Single-cell libraries are usually sequenced deeply enough to detect most genes.
Scientists often decide whether to sequence many cells with less detail or fewer cells with more detail. Sequencing more cells helps find the main types of cell populations. However, less detailed sequencing might not always provide enough information about specific genes.
Scientists use quality checks to decide how many cells to analyze. These checks look at things like the number of RNA counts, the number of genes found, and the amount of RNA from mitochondria. This helps them understand the true signals from the cells.
Learn More
- DNA sequencing
- Human Cell Atlas
- Disease Cell Atlas
- Single-cell analysis
- Single cell epigenomics
- Single-cell transcriptomics
- Single-cell multi-omics integration
- Tcr-seq
- Whole genome sequencing