AlphaFold facts for kids
AlphaFold is a special computer program that uses artificial intelligence (AI) to figure out the shapes of proteins. It was created by a company called DeepMind, which is part of Alphabet. Knowing a protein's shape helps scientists understand what it does in our bodies and in nature.
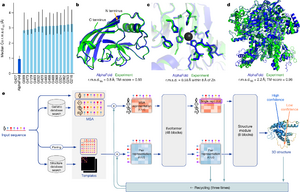
The first version, AlphaFold 1, won a big competition called CASP in 2018. It was really good at predicting the shapes of proteins that were very hard to figure out.
Then came AlphaFold 2 in 2020, and it did even better in the CASP competition. It was much more accurate than any other program. For many proteins, its predictions were almost perfect, scoring over 90 out of 100 on a test called GDT. This was a huge step forward for science. In 2021, the details of AlphaFold 2 were shared with the world, along with a free database of protein shapes.
On May 8, 2024, AlphaFold 3 was announced. This new version can predict not just single protein shapes, but also how proteins connect with other important molecules like DNA, RNA, and different small chemicals. It's much better at predicting these complex connections.
Because of their amazing work on AlphaFold, Demis Hassabis and John Jumper from Google DeepMind shared half of the 2024 Nobel Prize in Chemistry.
Contents
What Are Proteins and Why Do Their Shapes Matter?
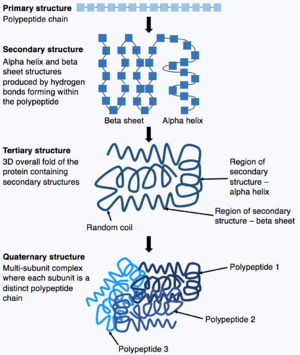
Proteins are like tiny machines inside all living things. They are made of long chains of smaller units called amino acids. These chains fold up into very specific three-dimensional (3-D) shapes. A protein's shape is super important because it determines what job the protein can do. For example, some proteins help us digest food, while others fight off sickness.
Scientists can find out protein shapes using special lab methods like X-ray crystallography. But these methods are expensive and take a lot of time. Over the last 60 years, scientists have found the shapes of about 170,000 proteins. However, there are over 200 million known proteins in all life forms! This is why computer programs like AlphaFold are so helpful.
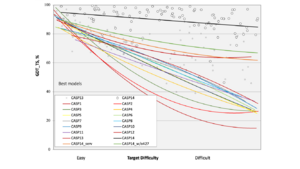
For many years, researchers have tried to use computers to guess protein shapes from their amino acid sequences. The CASP competition started in 1994 to challenge scientists to make the best predictions. Before AlphaFold, even the best programs struggled with difficult proteins. AlphaFold changed this by using artificial intelligence and deep learning.
How AlphaFold Works
DeepMind trained AlphaFold using information from over 170,000 proteins whose shapes were already known. The program uses a special AI technique called an "attention network." This is like how you might solve a jigsaw puzzle: you first connect small groups of pieces, and then you figure out how to join those groups into a bigger picture. AlphaFold does something similar, focusing on different parts of the protein to build the whole shape.
AlphaFold 1 (2018)
The first AlphaFold program, released in 2018, looked at huge databases of related DNA sequences. It tried to find connections between amino acids that were far apart in the protein chain but might be close together in the final 3-D shape. AlphaFold 1 was able to guess the distances between these amino acids, which helped it predict the protein's overall structure.
AlphaFold 2 (2020)
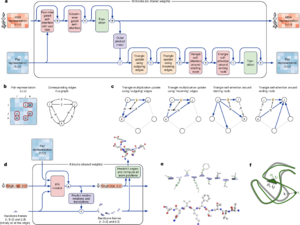
The 2020 version, AlphaFold 2, was very different and much more powerful than AlphaFold 1. Instead of separate steps, AlphaFold 2 used one big, connected system of AI networks. This system learned to recognize patterns in protein data in a very smart way.
A key part of AlphaFold 2 is how it refines its predictions. It's like it keeps improving its guess over and over again. Imagine it guessing a shape, then checking for errors, and then adjusting the shape to fix those errors. It does this many times until the predicted shape is very accurate and physically correct.
In October 2021, DeepMind updated AlphaFold 2 to predict not just single proteins, but also how proteins connect with each other to form "protein complexes." This was a big step, as many proteins work together in groups.
AlphaFold 3 (2024)
AlphaFold 3 was developed by Google DeepMind and Isomorphic Labs. It was announced on May 8, 2024. This version is a huge leap because it can predict the shapes of proteins when they are connected to other important molecules. This includes DNA, RNA, and various small chemicals called ligands and ions.
AlphaFold 3 uses a new AI design called "Pairformer." It also uses a "diffusion model," which starts with a cloud of atoms and slowly brings them together to form the correct 3-D shape. This new method is much more accurate for predicting how proteins interact with other molecules.
A free online tool called the AlphaFold server was created to let scientists use AlphaFold 3 for their research.
Competitions and Achievements
CASP13 (2018)
In December 2018, DeepMind's AlphaFold 1 won the 13th CASP competition. It was especially good at predicting the shapes of proteins that were considered the most difficult. AlphaFold 1 scored much higher than other teams, showing that AI could make a big difference in protein prediction.
CASP14 (2020)
In November 2020, AlphaFold 2 won CASP14. It made the best predictions for almost all the proteins in the competition. Its accuracy was so high that it was compared to shapes found using expensive lab methods like X-ray crystallography. This was a truly amazing achievement. For example, AlphaFold 2 helped scientists figure out the shape of a difficult protein that had puzzled them for ten years!
CASP15 (2022)
DeepMind did not enter CASP15 in 2022. However, many of the teams that did compete used AlphaFold or tools based on AlphaFold, showing its widespread impact.
AlphaFold Protein Structure Database
| Content | |
|---|---|
| Data types captured |
protein structure prediction |
| Organisms | all UniProt proteomes |
| Contact | |
| Research center | EMBL-EBI |
| Access | |
| Website | https://www.alphafold.ebi.ac.uk/ |
| Download URL | yes |
| Tools | |
| Web | yes |
| Miscellaneous | |
| License | CC-BY 4.0 |
| Curation policy | automatic |
The AlphaFold Protein Structure Database was launched on July 22, 2021. It's a joint project between AlphaFold and EMBL-EBI. When it first started, it had predicted shapes for almost all human proteins and proteins from 20 other important organisms.
By July 28, 2022, the database had grown to include the shapes of about 200 million proteins from 1 million different species. This means it covers almost every known protein on Earth! This huge database is a fantastic resource for scientists around the world.
What AlphaFold Can and Cannot Do
Even though AlphaFold is incredibly powerful, it has some limits:
- The database mostly shows shapes of single protein chains, not always how they work together in groups.
- Some parts of the predicted shapes might not be as certain as others.
- AlphaFold is best when it can compare a new protein to similar ones it has already learned from. It might not work as well for completely new or artificial proteins.
- It's not always perfect at predicting how a protein might change its shape.
- AlphaFold 3 can predict interactions with some other molecules, but not all of them. For example, it doesn't fully predict all the sugar molecules that can attach to proteins.
- Sometimes, AlphaFold might create a shape that looks physically impossible, like a protein chain that ties itself in knots.
Real-World Uses
AlphaFold has already been used to predict the shapes of proteins from SARS-CoV-2, the virus that causes COVID-19. This helped scientists understand the virus better, even before lab experiments could confirm the shapes. For example, AlphaFold 2 accurately predicted the shape of a protein called ORF3a, which helps the virus escape from human cells and causes inflammation.
More to Explore
- Folding@home
- Foldit
- AlphaZero
- AlphaGo
- AlphaGeometry
See also
 In Spanish: AlphaFold para niños
In Spanish: AlphaFold para niños